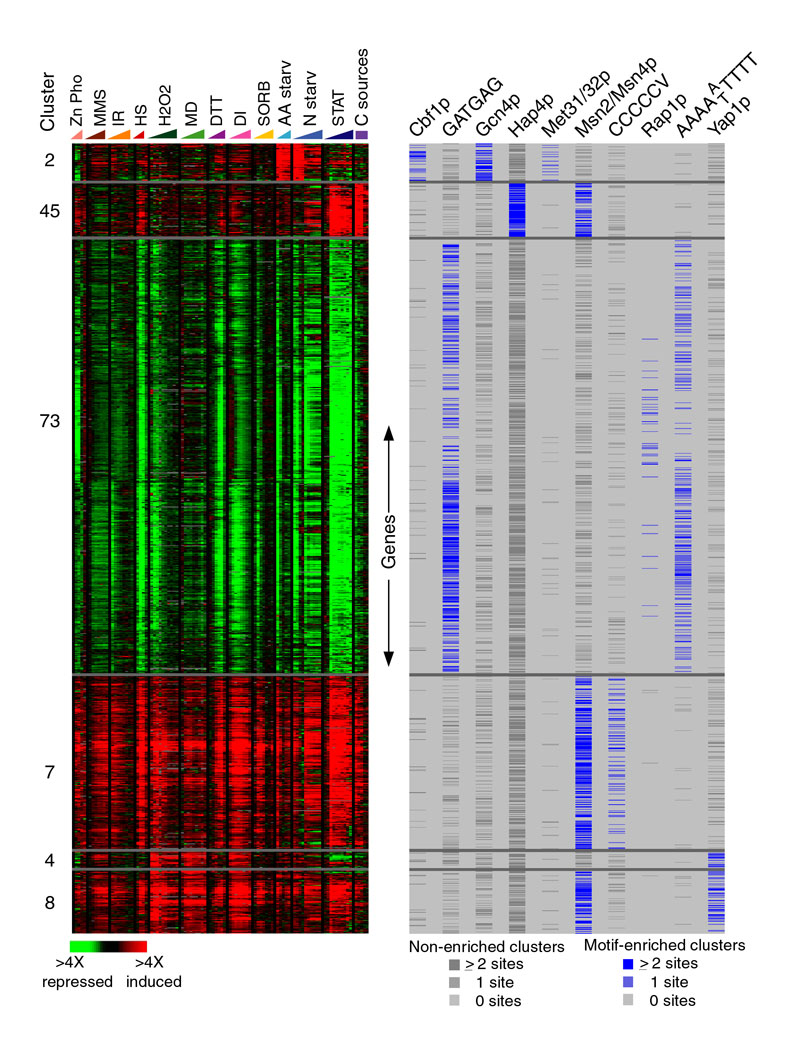
The enrichment of each cluster for genes that contain known transcription factor binding sites in their promoters was measured based on the hypergeometric distribution, as described in Materials and Methods. The gene expression data are shown (left panel) as described in Figure 1 for genes that were assigned to cluster 2 (amino acid biosynthesis genes), cluster 45 (respiration genes) cluster 73 (genes repressed as part of the environmental stress response (ESR)), cluster 7 (genes induced as part of the ESR), cluster 4 (oxidative stress defense genes), cluster 8 (genes conditionally regulated by Yap1p or Msn2/Msn4p). The genes were assigned to each cluster with a membership cutoff of 0.08, with the exception of cluster 2 for which a cutoff of 0.06 was used. The hypergeometric distribution was used to measure the statistical enrichment of promoters containing the binding sites of Cbf1p (TGACGTG), the ESR motif GATGAG, the binding site of Hap2/3/4p (CCAAT), Met31/32p (AAACTGTG), Msn2/Msn4p (CCCCT), a C-rich element identified in cluster 7 (CCCCCV where V is any nucleotide but T), Rap1p (ACACCCAYACAY where Y is C or T), the ESR motif AAAAWTTTT (where W is A or T), and Yap1p (TTAGTMA where M is C or A). For each gene displayed in the left panel, the copy number of the denoted transcription factor binding sites in the gene's promoter is indicated by a colored box (right panel): the copy number is indicated with a blue box only if the cluster to which the gene belonged was statistically enriched (P value < 2x10-4) for the indicated binding site, whereas the copy number is indicated with a dark gray box if the cluster to which the gene belonged was not statistically enriched for the site. Download the complete results.